Our technology
Upload your genomic sequences. Our servers take care of the complete analysis no setup, no local servers, no hassle.
gen4epi is a cloud-based bioinformatics platform that automates the entire genomic analysis workflow, from uploading and processing genomic sequences to generating complete bioinformatics results on our servers. Once generated, an artificial intelligence–based assistant supports the interpretation of results, helping users understand patterns, genetic relationships, and epidemiological contexts without requiring advanced computing skills.
Sequences are stored and compressed using an optimized algorithm that preserves all genetic information while minimizing the required storage space. Our servers handle the processing remotely, without using the user’s local resources, ensuring speed, scalability, and security.
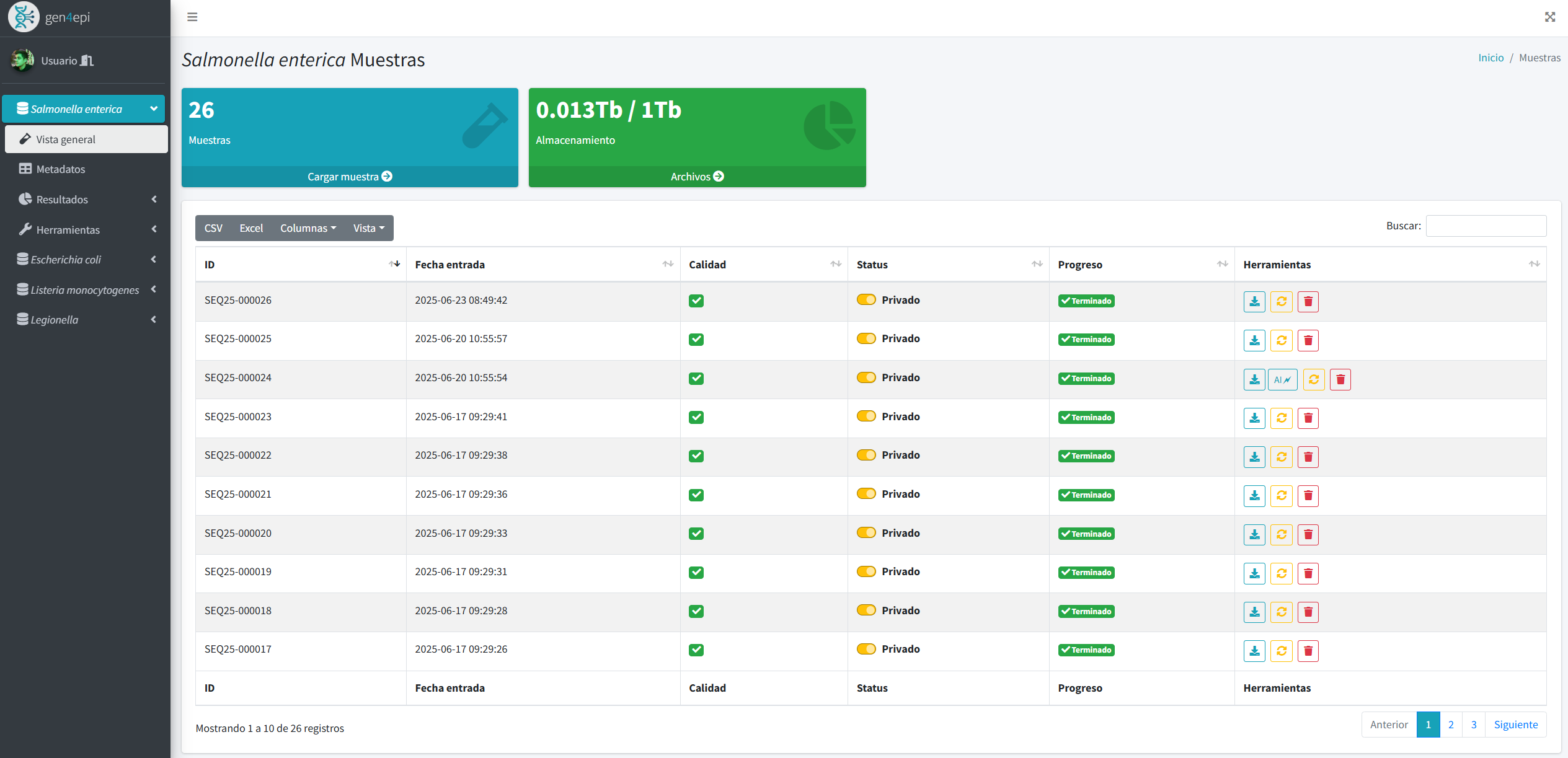
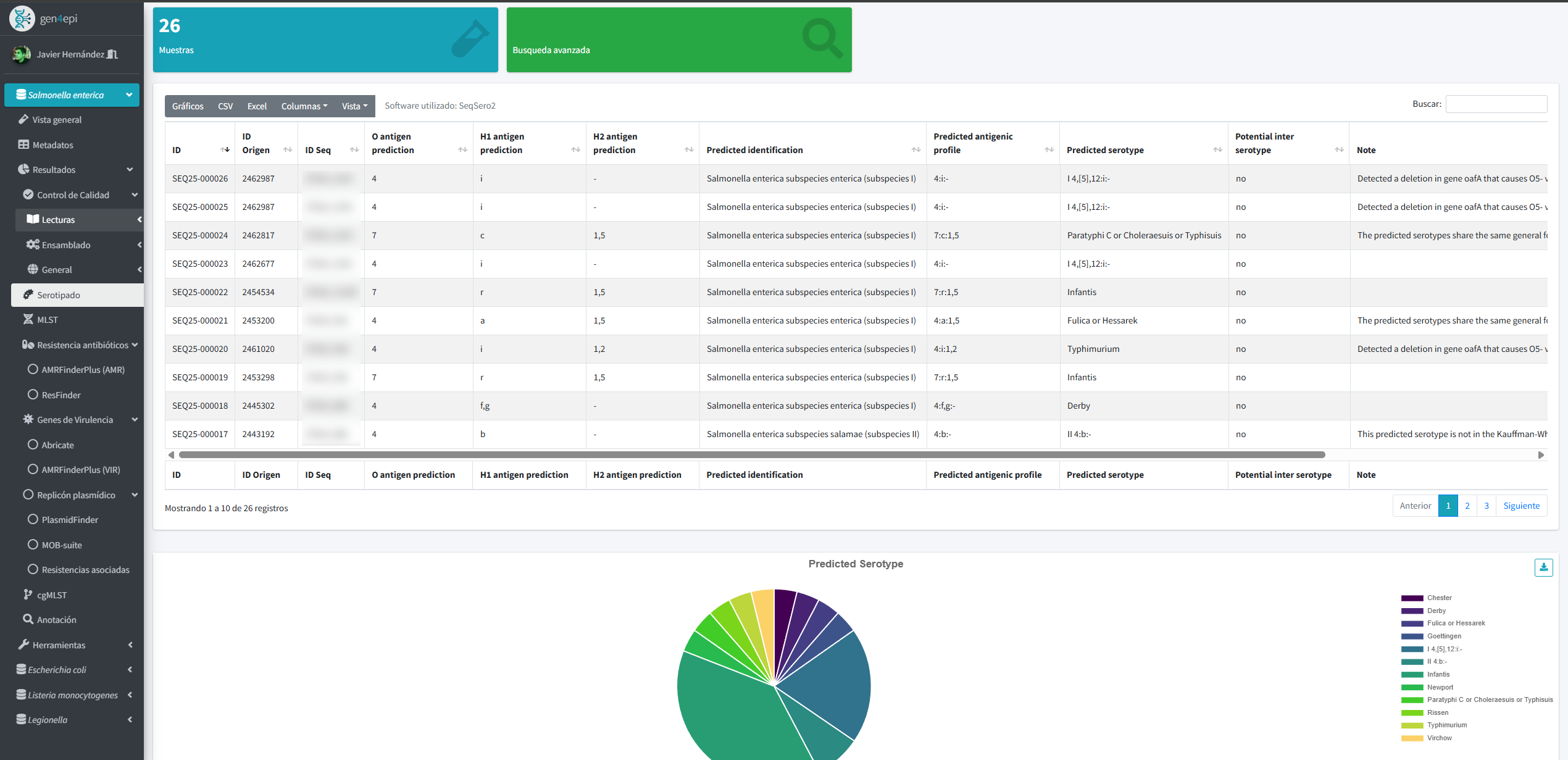
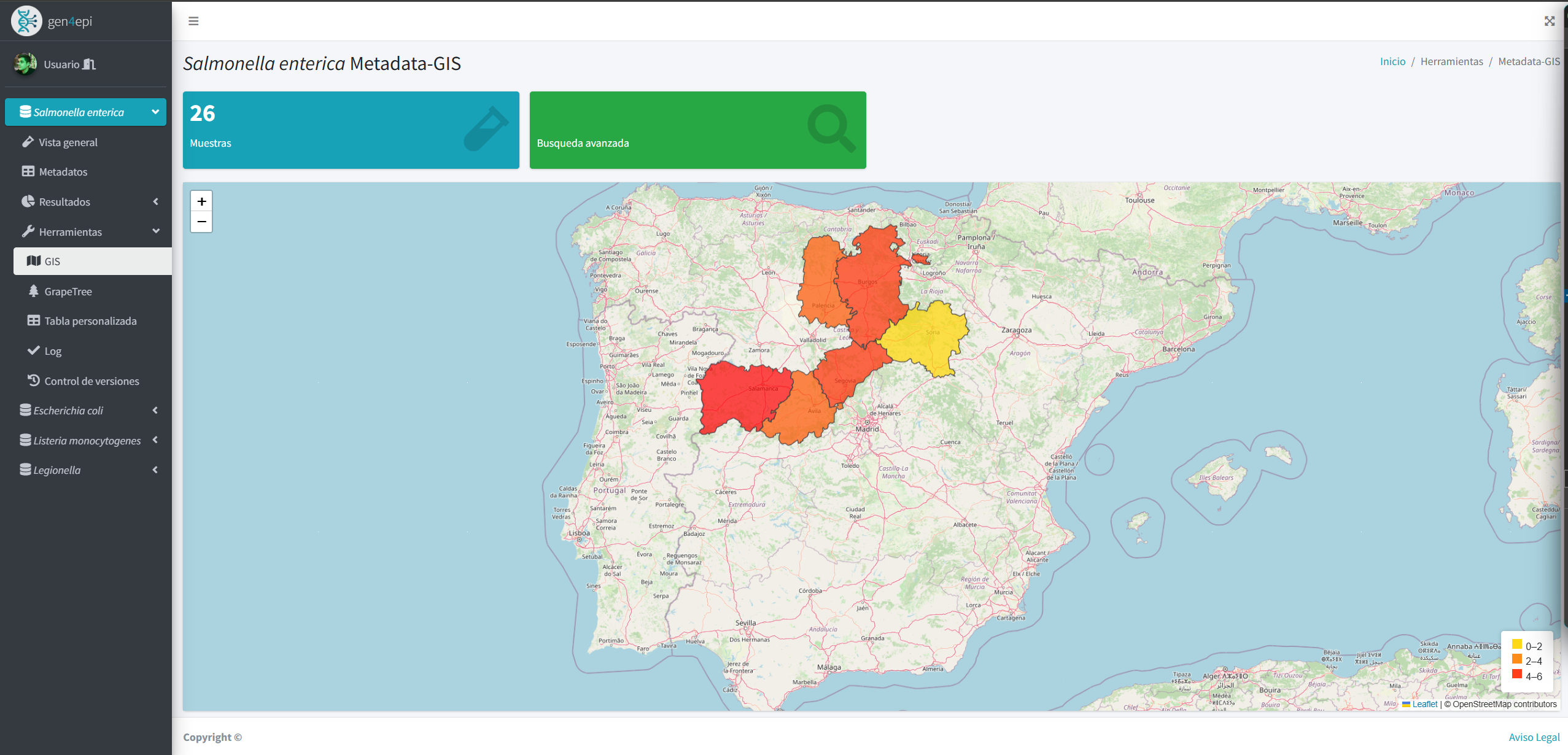
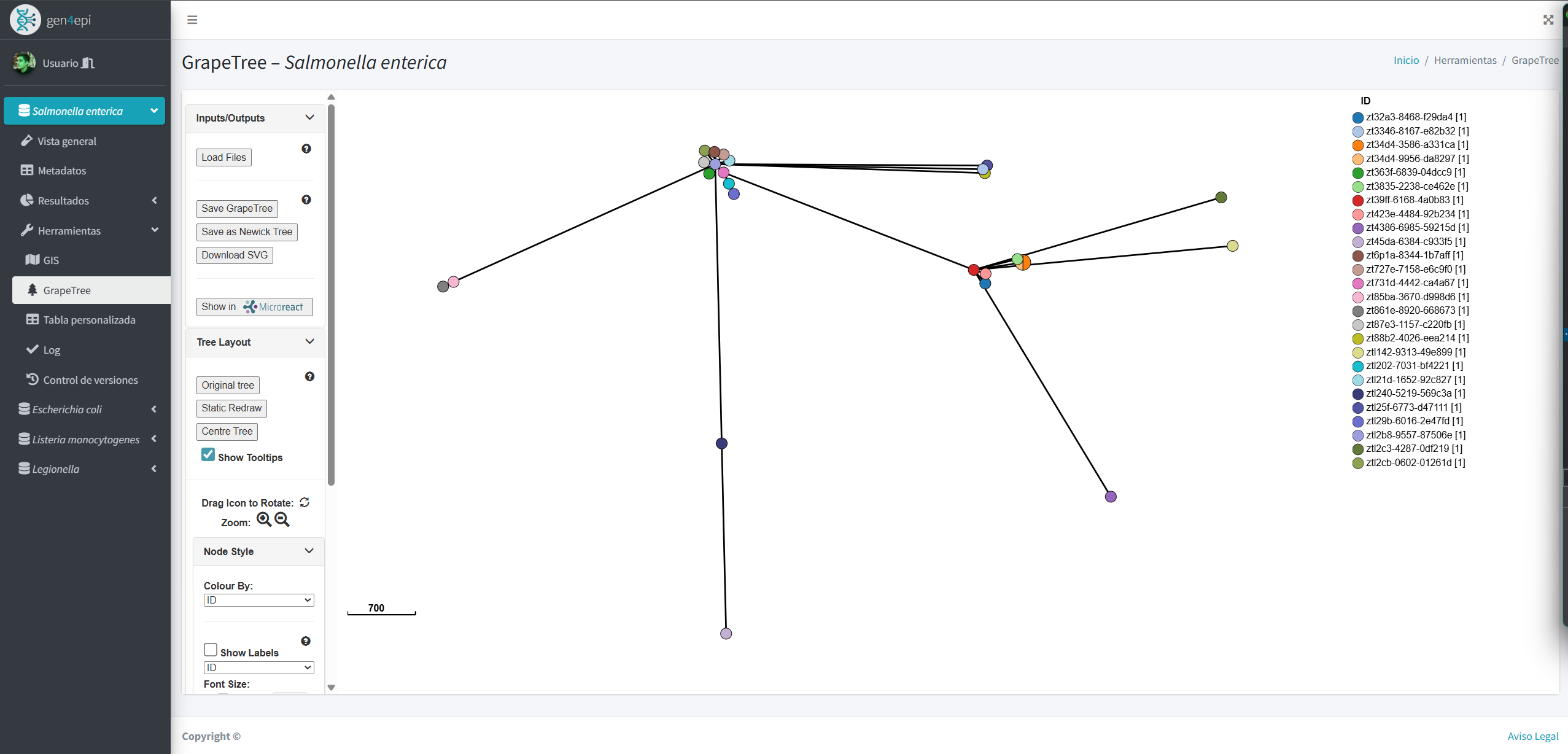
Results include dynamic tables, interactive visualizations, and automated reports that make it easier to understand large volumes of data. In addition, the system integrates GIS tools and phylogenetic analyses to explore epidemiological patterns that reflect the genetic evolution of strains.
With its integrated artificial intelligence, gen4epi detects relevant patterns, generates early warnings for unusual behavior, and contributes to faster, more accurate, and more proactive epidemiological surveillance.